trc shrna library (Broad Clinical Labs)
Structured Review

Trc Shrna Library, supplied by Broad Clinical Labs, used in various techniques. Bioz Stars score: 96/100, based on 693 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more
https://www.bioz.com/result/trc shrna library/product/Broad Clinical Labs
Average 96 stars, based on 693 article reviews
Images
1) Product Images from "CDKN1B (p27 kip1 ) enhances drug tolerant persister CTCs by restricting polyploidy following mitotic inhibitors"
Article Title: CDKN1B (p27 kip1 ) enhances drug tolerant persister CTCs by restricting polyploidy following mitotic inhibitors
Journal: bioRxiv
doi: 10.1101/2024.02.20.581202

Figure Legend Snippet: 4A: DTX treatment upregulates CDKN1B expression in a fraction of CTCs. Representative scatter plot showing untreated (NT) and DTX-treated (10nM, day 7) CTCs, stained with antibody against CDKN1B and analyzed by flow cytometry. 4B: Bar graph: (mean ± SEM) increase in CDKN1B-positive cells in DTX-treated per pro CTCs (BRx-68, BRx-82, BRx-142, BRx-211, BRx-250, BRx-292), compared with untreated cells (NT). n=3 biological repeats, with p-value calculated using two-tailed Student’s T-test (** p<0.01). 4C: Suppression of DTPs in per pro CTCs following CDKN1B knockdown using transient siRNA transfection. CTC cultures were transfected with siCDKN1B (red) or siControl (blue), and after three days treated with increasing concentrations of DTX (Knockdown efficiency is show in Supplemetary Figure S4F). Knockdown of CDKN1B decreases the minimal residual fraction of viable CTCs in per pro CTC cultures (BRx-68, BRx-82, BRx-142), but it has no effect in the already low viable cell fraction in per def CTC cultures (BRx07, BRx-29, BRx-394). 1-E max at each DTX dose was measured (n= 3 biological repeats). p-value (p<0.0001 for each of the BRx-68, BRx-82 and BRx-142 CTCs) was calculated using a two-tailed Student’s T-Test. 4D : Stable CDKN1B mRNA knockdown by two different shRNA constructs (sh1B-H3, sh1B-H4), shown using bar graph (mean ± SEM) in per pos CTCs (BRx-82, BRx-142), compared with scrambled control. P-value calculated using two-tailed Student’s T-Test (**** p<0.0001). 4E: CDKN1B knockdown suppresses regrowth of DTPs following pulse DTX treatment of per pro CTCs (BRx-82, BRx-142). Infection of CTC cultures with either sh1B-H3 (red) or sh1B-H4 (green), followed by DTX exposure (10nM, 16 hrs) abolishes the regrowth observed with shScrambled control (blue). A representative graph, from at least two biological repeats, is shown. 4F: CDKN1B knockdown efficiency following siRNA transfection of CTC cultures (BRx-142) is represented by bar graph (mean ± SEM). p-value was calculated using two-tailed Student’s T-Test (*** p<0.01). 4G: Bar graphs (mean ± SEM) showing the quantification of mitotic (pHH3-positive) 4N and ≥8N BRx-142 CTCs, shown for untreated (NT, blue) and DTX-treated (red; 10nM, day 7) cultures, following transfection with either siCDKN1B or siControl. CDKN1B knockdown increases the number of mitotic >8N CTCs following pulse DTX, but not the number of mitotic 4N CTCs. P-value calculated using two-tailed Student’s T-Test (** p<0.01, ***p<0.001). n=3 biological triplicates.
Techniques Used: Expressing, Staining, Flow Cytometry, Two Tailed Test, Knockdown, Transfection, shRNA, Construct, Control, Infection
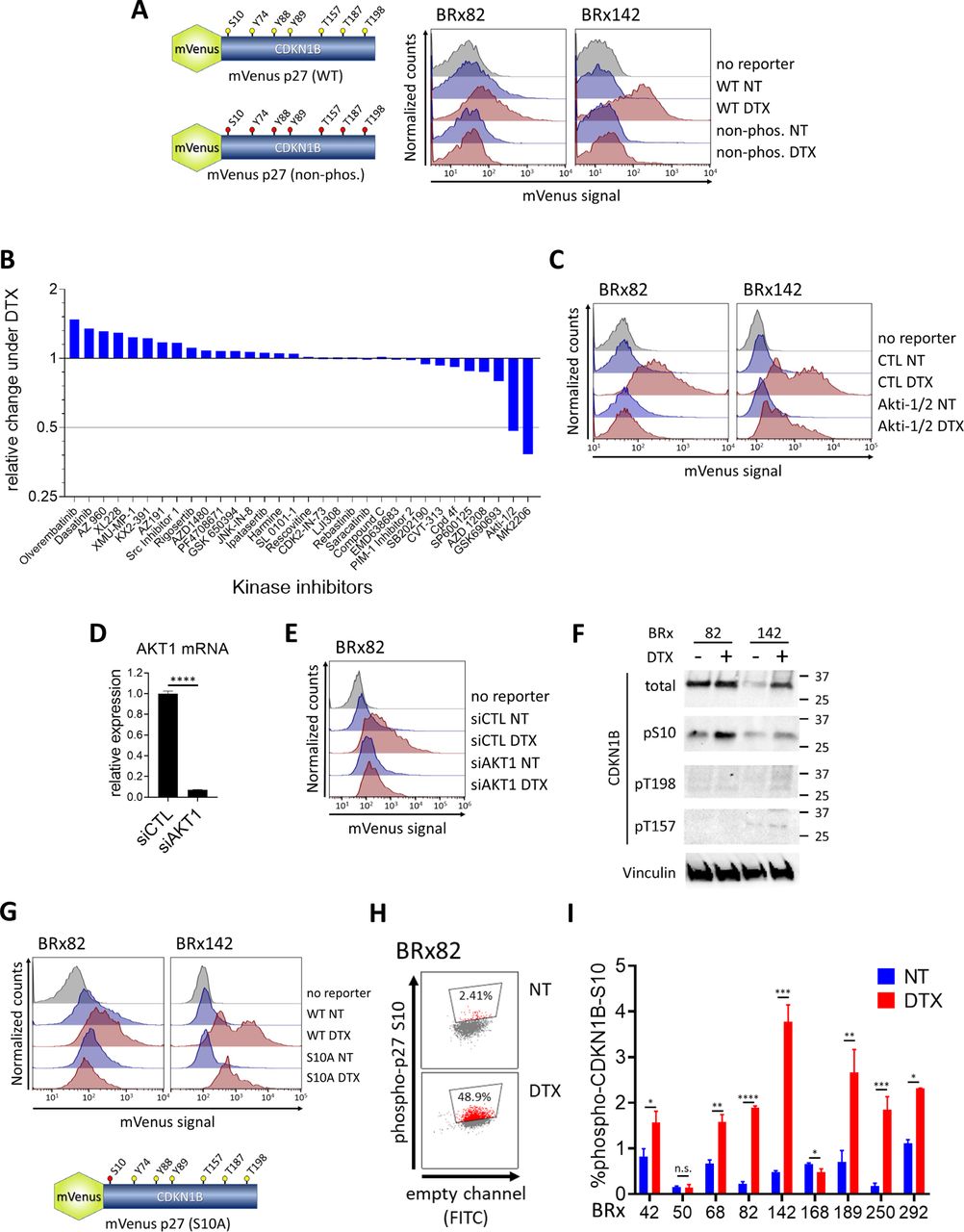
Figure Legend Snippet: 5A: Stabilization of mVenus-p27 reporter construct following DTX exposure. Left: schema of the reporter, fusing a fluorescent tag (mVenus) to the catalytically inactive CDKN1B coding sequence of CDKN1B, driven by the CMV promoter. Phosphorylation target residues are shown in the wild-type (WT) reporter, all of which are mutated to alanine in the mutant construct (non-phos.) Right: Increased mVenus fluorescence signal (flow cytometry) following treatment of CTC cultures (BRx-82, BRx-142) with DTX (10nM, 4 days), indicating stabilization of the reporter protein. Mutation of all 7 candidate phosphorylation target residues abrogates stabilization of the reporter. One of two biological repeats is shown. 5B : Waterfall plot, showing a kinase inhibitor screen to identify candidates mediating stabilization of the mVenus-p27 reporter, following treatment of CTCs (BRx-142) with DTX (10nM, 4 days). Among 32 kinase inhibitors targeting 15 kinases previously implicated in stabilizing CDKN1B, two compounds targeting AKT (Akti-1/2 and MK2206) suppress DTX-mediated stabilization of CDKN1B. Bar graphs represent the ratio of the percent reporter-positive cells exposed to DTX along with inhibitor as a function of the percent reporter-positive cells exposed to DTX alone. 5C: Suppression of CDKN1B reporter stabilization in CTC cultures (BRx-82, BRx-142) treated with DTX (10nM, 4 days) in the presence of the AKT inhibitor Akti-1/2 (100mM). mVenus fluorescence signal in live cells was measured using flow cytometry, with one of two biological repeats shown. 5D: Knockdown of the AKT1 isoform using siRNA transfection in CTC cultures (BRx-82) stably expressing the CDKN1B reporter. Bar graph (mean ± SEM), with p-value was calculated using two-tailed Student’s T-Test (**** p<0.0001 5E: Suppression of CDKN1B reporter stabilization in DTX-treated CTCs (BRx-82) following transfection with si AKT1 , compared with siControl. Shown is one of two biological repeats. 5F: Western blot analysis showing increased phosphorylation of one of the three AKT phosphorylation sites on native CDKN1B protein, serine-10 (S10), in CTC cultures (BRX-82, BRx-142) following treatment with DTX (10nM, 4 days). The other two AKT mediated phosphorylation sites (T198 and T157) are less evident by Western blotting . Total CDKN1B protein levels are shown (with and without DTX exposure), along with Vinculin expression (loading control). 5G: Mutation of the CDKN1B S10 residue within the mVenus reporter is sufficient to reduce DTX-mediated protein stabilization. The position of the serine 10 residue (mutated to alanine) is shown in the schema (mVenus p27 (S10A). Mutation of S10 reduces CDKN1B stabilization in CTCs (BRx-82, BRx-142) following DTX exposure (10nM, 4 days), compared with wild type (WT) reporter. mVenus fluorescence signal in live cells measured using flow cytometry, with one of two biological repeats shown. 5H: Flow cytometric analysis, showing increased subpopulation of CTC cultures (BRx-82) with detectable endogenous phospho-serine 10 (S10)-CDKN1B protein, following DTX exposure, compared with untreated cultures (NT). A representative scatter plot is shown. 5I: Bar graph, showing quantification of the fraction of per pro CTC cultures (BRx-42, BRx-50, BRx-68, BRx-82, BRx-142, BRx-168, BRx-189, BRx-292) with phosphorylated endogenous Serine 10 (S10)-CDKN1B. Percent S10-CDKN1B positive cells (mean ± SEM) following DTX exposure or untreated (NT) is shown, with p-value calculated using two-tailed Student’s T-Test (* p<0.05, ** p<0.01, *** p<0.001, **** p<0.0001).
Techniques Used: Construct, Sequencing, Phospho-proteomics, Mutagenesis, Fluorescence, Flow Cytometry, Knockdown, Transfection, Stable Transfection, Expressing, Two Tailed Test, Western Blot, Control, Residue


